pMELTS¶
Versions of MELTS implemented are:
- MELTS v. 1.0.2 ➞ (rhyolite-MELTS, Gualda et al., 2012)
- MELTS v. 1.1.0 ➞ (rhyolite-MELTS + new CO2, works at the ternary
minimum)
- MELTS v. 1.2.0 ➞ (rhyolite-MELTS + new H2O + new CO2)
- pMELTS v. 5.6.1
Initialize tools and packages that are required to execute this notebook.¶
from thermoengine import equilibrate
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
Create a pMELTS v 5.6.1 instance.¶
Rhyolite-MELTS version 1.0.2 is the default model.
melts = equilibrate.MELTSmodel(version="5.6.1")
Optional: Generate some information about the implemented model.¶
oxides = melts.get_oxide_names()
phases = melts.get_phase_names()
#print (oxides)
#print (phases)
Required: Input initial composition of the system (liquid), in wt% or grams of oxides.¶
Early Bishop Tuff average melt inlusion composition
feasible = melts.set_bulk_composition({'SiO2': 45.47,
'TiO2': 0.11,
'Al2O3': 4.0,
'Fe2O3': 0.585,
'Cr2O3': 0.0,
'FeO': 6.696,
'MnO': 0.0,
'MgO': 38.53,
'NiO': 0.0,
'CoO': 0.0,
'CaO': 3.59,
'Na2O': 0.31,
'K2O': 0.0,
'P2O5': 0.0,
'H2O': 0.0})
Optional: Suppress phases that are not required in the simulation.¶
b = melts.get_phase_inclusion_status()
melts.set_phase_inclusion_status({'Actinolite':False, 'Aegirine':False, \
'Aenigmatite':False, 'Akermanite':False, 'Andalusite':False, \
'Anthophyllite':False, 'Apatite':False, 'Biotite':False, 'Chromite':False, \
'Coesite':False, 'Corundum':False, 'Cristobalite':False, 'Cummingtonite':False, \
'Fayalite':False, 'Forsterite':False, 'Gehlenite':False, 'Hematite':False, \
'Hornblende':False, 'Ilmenite':False, 'Ilmenite ss':False, 'Kalsilite':False, \
'Kalsilite ss':False, 'Kyanite':False, 'Leucite':False, 'Lime':False, \
'Liquid Alloy':False, 'Magnetite':False, 'Melilite':False, 'Muscovite':False, \
'Nepheline':False, 'Nepheline ss':False, 'OrthoOxide':False, 'Panunzite':False, \
'Periclase':False, 'Perovskite':False, 'Phlogopite':False, 'Quartz':False, \
'Rutile':False, 'Sanidine':False, 'Sillimanite':False, 'Solid Alloy':False, \
'Sphene':False, 'Tridymite':False, 'Whitlockite':False})
a = melts.get_phase_inclusion_status()
for phase in b.keys():
if b[phase] != a[phase]:
print ("{0:<15s} Before: {1:<5s} After: {2:<5s}".format(phase, repr(b[phase]), repr(a[phase])))
Actinolite Before: True After: False
Aegirine Before: True After: False
Aenigmatite Before: True After: False
Akermanite Before: True After: False
Andalusite Before: True After: False
Anthophyllite Before: True After: False
Apatite Before: True After: False
Biotite Before: True After: False
Chromite Before: True After: False
Coesite Before: True After: False
Corundum Before: True After: False
Cristobalite Before: True After: False
Cummingtonite Before: True After: False
Fayalite Before: True After: False
Forsterite Before: True After: False
Gehlenite Before: True After: False
Hematite Before: True After: False
Hornblende Before: True After: False
Ilmenite Before: True After: False
Ilmenite ss Before: True After: False
Kalsilite Before: True After: False
Kalsilite ss Before: True After: False
Kyanite Before: True After: False
Leucite Before: True After: False
Lime Before: True After: False
Liquid Alloy Before: True After: False
Magnetite Before: True After: False
Melilite Before: True After: False
Muscovite Before: True After: False
Nepheline Before: True After: False
Nepheline ss Before: True After: False
OrthoOxide Before: True After: False
Panunzite Before: True After: False
Periclase Before: True After: False
Perovskite Before: True After: False
Phlogopite Before: True After: False
Quartz Before: True After: False
Rutile Before: True After: False
Sanidine Before: True After: False
Sillimanite Before: True After: False
Solid Alloy Before: True After: False
Sphene Before: True After: False
Tridymite Before: True After: False
Whitlockite Before: True After: False
Compute the equilibrium state at some specified T (°C) and P (MPa).¶
Print status of the calculation.
output = melts.equilibrate_tp(1300.0, 1000.0, initialize=True)
(status, t, p, xmlout) = output[0]
print (status, t, p)
success, Optimal residual norm. 1300.0 1000.0
Summary output of equilibrium state …¶
melts.output_summary(xmlout)
T (°C) 1300.00
P (MPa) 1000.00
Augite 15.6279 (g) Na0.05Ca0.67Fe''0.10Mg0.97Fe'''0.04Ti0.01Al0.32Si1.84O6
Olivine 53.9240 (g) (Ca0.00Mg0.91Fe''0.09Mn0.00Co0.00Ni0.00)2SiO4
Spinel 1.1339 (g) Fe''0.12Mg0.88Fe'''0.10Al1.89Cr0.00Ti0.00O4
Liquid 4.0410 (g) wt%:SiO2 48.94 TiO2 0.48 Al2O3 19.05 Fe2O3 1.96 Cr2O3 0.00 FeO 5.04 MnO 0.00 MgO
11.73 NiO 0.00 CoO 0.00 CaO 8.39 Na2O 4.42 K2O 0.00 P2O5 0.00 H2O 0.00
Orthopyroxene 24.5641 (g) Na0.01Ca0.06Fe''0.15Mg1.67Fe'''0.02Ti0.00Al0.22Si1.88O6
Compute the equilibrium state at some specified S (J/K) and P (MPa).¶
s = melts.get_property_of_phase(xmlout,'System', 'Entropy')
print ("{0:<20s} {1:13.6e} {2:<10s}".format('Entropy', s, melts.get_units_of_property('Entropy')))
output = melts.equilibrate_sp(s, p, initialize=True)
(status, t, p, xmlout) = output[0]
melts.output_summary(xmlout)
Entropy 2.505416e+02 J/K
T (°C) 1300.00
P (MPa) 1000.00
Augite 15.6279 (g) Na0.05Ca0.67Fe''0.10Mg0.97Fe'''0.04Ti0.01Al0.32Si1.84O6
Olivine 53.9240 (g) (Ca0.00Mg0.91Fe''0.09Mn0.00Co0.00Ni0.00)2SiO4
Spinel 1.1339 (g) Fe''0.12Mg0.88Fe'''0.10Al1.89Cr0.00Ti0.00O4
Liquid 4.0410 (g) wt%:SiO2 48.94 TiO2 0.48 Al2O3 19.05 Fe2O3 1.96 Cr2O3 0.00 FeO 5.04 MnO 0.00 MgO
11.73 NiO 0.00 CoO 0.00 CaO 8.39 Na2O 4.42 K2O 0.00 P2O5 0.00 H2O 0.00
Orthopyroxene 24.5641 (g) Na0.01Ca0.06Fe''0.15Mg1.67Fe'''0.02Ti0.00Al0.22Si1.88O6
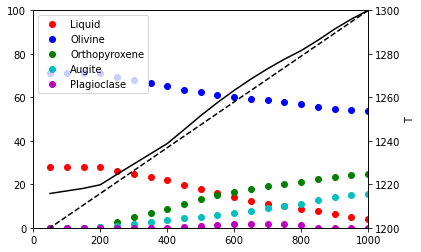
Run the sequence of calculations along a S=constant, P path:¶
Output is sent to an Excel file and plotted in the notebook
number_of_steps = 20
s_increment_of_steps = 0.0
p_increment_of_steps = -50.0
plotPhases = ['Liquid', 'Olivine', 'Orthopyroxene', 'Augite', 'Plagioclase']
# matplotlib colors b : blue, g : green, r : red, c : cyan, m : magenta, y : yellow, k : black, w : white.
plotColors = [ 'ro', 'bo', 'go', 'co', 'mo']
wb = melts.start_excel_workbook_with_sheet_name(sheetName="Summary")
melts.update_excel_workbook(wb, xmlout)
n = len(plotPhases)
xPlot = np.zeros(number_of_steps)
yPlot = np.zeros((n, number_of_steps))
y2Plot = np.linspace(t, t-100.0, number_of_steps)
y3Plot = np.linspace(t, t-100.0, number_of_steps)
xPlot[0] = p
for i in range (0, n):
yPlot[i][0] = melts.get_property_of_phase(xmlout, plotPhases[i])
y2Plot[0] = t
plt.ion()
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_xlim([min(p, p+p_increment_of_steps*number_of_steps), max(p, p+p_increment_of_steps*number_of_steps)])
ax.set_ylim([0., 100.])
ax2 = ax.twinx()
ax2.set_ylabel('T', color='k')
ax2.set_ylim([t-100, t])
graphs = []
for i in range (0, n):
graphs.append(ax.plot(xPlot, yPlot[i], plotColors[i]))
graphs.append(ax2.plot(xPlot, y2Plot, 'k-'))
graphs.append(ax2.plot(xPlot, y3Plot, 'k--'))
handle = []
for (graph,) in graphs:
handle.append(graph)
ax.legend(handle, plotPhases, loc='upper left')
for i in range (1, number_of_steps):
output = melts.equilibrate_sp(s+s_increment_of_steps, p+p_increment_of_steps)
(status, t, p, xmlout) = output[0]
print ("{0:<30s} {1:8.2f} {2:8.2f}".format(status, t, p))
xPlot[i] = p
for j in range (0, n):
yPlot[j][i] = melts.get_property_of_phase(xmlout, plotPhases[j])
y2Plot[i] = t
j = 0
for (graph,) in graphs:
graph.set_xdata(xPlot)
if j < n:
graph.set_ydata(yPlot[j])
elif j == n:
graph.set_ydata(y2Plot)
j = j + 1
fig.canvas.draw()
melts.update_excel_workbook(wb, xmlout)
melts.write_excel_workbook(wb, "MELTSv102summary.xlsx")
success, Optimal residual norm. 1295.97 950.00
success, Optimal residual norm. 1291.41 900.00
success, Optimal residual norm. 1286.25 850.00
success, Optimal residual norm. 1281.49 800.00
success, Optimal residual norm. 1277.55 750.00
success, Optimal residual norm. 1273.18 700.00
success, Optimal residual norm. 1268.41 650.00
success, Optimal residual norm. 1263.21 600.00
success, Optimal residual norm. 1257.61 550.00
success, Optimal residual norm. 1251.57 500.00
success, Optimal residual norm. 1245.10 450.00
success, Optimal residual norm. 1238.75 400.00
success, Optimal residual norm. 1234.03 350.00
success, Optimal residual norm. 1229.33 300.00
success, Optimal residual norm. 1224.60 250.00
success, Optimal residual norm. 1219.79 200.00
success, Optimal residual norm. 1218.15 150.00
success, Optimal residual norm. 1216.97 100.00
success, Optimal residual norm. 1215.78 50.00